|
When sufficient unique identifying indexes are available, many samples, including samples from multiple projects, can be pooled and sequenced on higher throughput platforms which minimizes costs for all samples in the pool. It is well known that identifying DNA sequences (commonly called indexes, tags, or barcodes we use the term “indexes” throughout) can be incorporated during sample preparation for NGS (i.e., library construction) so that multiple samples can be pooled prior to NGS, thereby allowing the sequencing costs to be divided among the samples (see Faircloth & Glenn, 2012 and references therein). This creates a fundamental challenge: how do we efficiently create and pool large numbers of samples so that we can divide the cost of high capacity NGS sequencing runs among many samples, thereby reducing the cost per sample? Frustratingly, within every NGS platform, the lowest-cost sequencing kits have the highest costs per read ( Glenn, 2011 Glenn, 2016). Thus, the large number of reads from a typical NGS run comes with a relatively large buy-in cost but yields an extremely low cost per read. Substantial progress has been made in developing new, lower-cost instruments, but much less progress has been made in reducing the cost of sequencing runs (cf., Glenn, 2011 vs. Next-generation DNA sequencing (NGS) has facilitated a wide variety of benefits in the life sciences ( Ansorge, 2009 Goodwin, McPherson & McCombie, 2016), and NGS instruments have an ever-growing capacity to generate more reads per run. A fast and easy to use program to demultiplex amplicon pools with internal indexes is available at. A simple web page to design fusion primers compatible with iTru primers is available at. Thus, we show that these methods facilitate amplicon library construction for Illumina instruments at reduced cost with increased flexibility. We demonstrate the utility and versatility of our approaches with results from six projects using different implementations of our protocols. 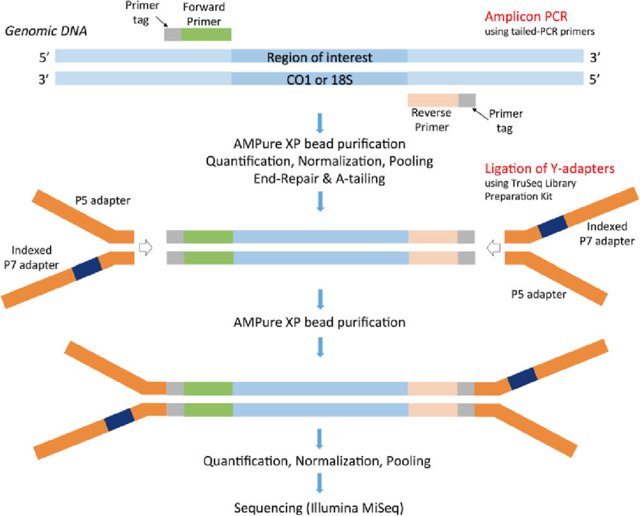
The number of sequencing reads from the amplicon pools can be adjusted, facilitating deep sequencing when required or reducing sequencing costs per sample to an economically trivial amount when deep coverage is not needed. The resulting quadruple-indexed amplicons have diversity at most base positions and can be pooled with any standard Illumina library for sequencing. The initial PCR products then serve as template for a second round of PCR with dual-indexed iTru or iNext primers (also used combinatorially) to make full-length libraries. These fusion primers can be used combinatorially to amplify samples within a 96-well plate (8 forward primers + 12 reverse primers yield 8 × 12 = 96 combinations), and the resulting amplicons can be pooled. In the variant we use most frequently for large-scale projects, we fuse partial adapter sequences (TruSeq or Nextera) onto the 5′ end of locus-specific PCR primers with variable-length tag sequences between the adapter and locus-specific sequences. Here, we describe multiple ways to use the Adapterama system and other approaches for amplicon sequencing on Illumina instruments. In Adapterama I, we presented universal stubs and primers to produce thousands of unique index combinations and a modifiable system for incorporating them into Illumina libraries. In many cases, NGS amplicon sequencing remains overly expensive and inflexible, with library preparation strategies relying upon the fusion of locus-specific primers to full-length adapter sequences with a single identifying sequence or ligating adapters onto PCR products. Next-generation sequencing (NGS) of amplicons is used in a wide variety of contexts. Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) PeerJ 7: e7786 
Cite this article Glenn TC, Pierson TW, Bayona-Vásquez NJ, Kieran TJ, Hoffberg SL, Thomas IV JC, Lefever DE, Finger JW, Gao B, Bian X, Louha S, Kolli RT, Bentley KE, Rushmore J, Wong K, Shaw TI, Rothrock Jr MJ, McKee AM, Guo TL, Mauricio R, Molina M, Cummings BS, Lash LH, Lu K, Gilbert GS, Hubbell SP, Faircloth BC. This work may be freely reproduced, distributed, transmitted, modified, built upon, or otherwise used by anyone for any lawful purpose. 29 Department of Environmental Sciences and Engineering, University of North Carolina, Chapel Hill, NC, United States of America DOI 10.7717/peerj.7786 Published Accepted Received Academic Editor Gerard Lazo Subject Areas Bioinformatics, Genetics, Genomics, Public Health Keywords MiSeq, Next generation sequencing, Quadruple indexing, Hierarchical indexing, Multiplexing, Fusion primers, Internal tagging, PCR, Libraries Licence This is an open access article, free of all copyright, made available under the Creative Commons Public Domain Dedication.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
